Coil Library Methodology
Below is a brief description of the methods used to generate the files in the coil library. A reference is forthcoming.
Secondary Structure Determination
Secondary structure in the Protein Coil Library is determined in a similar fashion as described by Srinivasan and Rose (1). Their method tiles Ramachandran space into "mesostate" regions, each assigned a unique identifier. A protein chain can be described as a sequence of mesostate identifiers. Secondary structure is determined from this sequence by identifying patterns common to α-helix, β-strand, and turns. A consequence of using this method to determine secondary structure is that only backbone torsions contribute to classification; no hydrogen bonding criteria is needed.
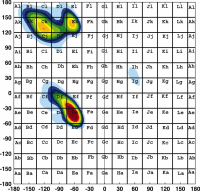 The method used to determine secondary structure in the Protein Coil
Library is also based on torsion-angle mesostate strings; however, the mesostate
regions used are smaller than in reference (1),
30 by 30 degree bins versus 60 by 60 degree bins. The bins are illustrated in the
figure at left, overlaid on to a contour plot of Ramachandran torsions calculated
by Hovmöller et. al. (2).
Given these definitions, secondary structure is determined using the match
hieirarchy below:
The method used to determine secondary structure in the Protein Coil
Library is also based on torsion-angle mesostate strings; however, the mesostate
regions used are smaller than in reference (1),
30 by 30 degree bins versus 60 by 60 degree bins. The bins are illustrated in the
figure at left, overlaid on to a contour plot of Ramachandran torsions calculated
by Hovmöller et. al. (2).
Given these definitions, secondary structure is determined using the match
hieirarchy below:
| SS Type | SS Code | Mesostates | Description |
| α-helix | H | De, Df, Ed, Ee, Ef, Fe | A region is identified as helix if there are five or more contiguous residues in the mesostate set. |
| β-strand | E | Bj, Bk, Bl, Cj, Ck, Cl, Dj, Dk, Dl | A region is defined as strand if there are three or more contiguous residues in the mesostate set. |
| β-turn | T | EfDf, EeEf, EfEf, EfDg, EeDg, EeEe, EfCg, EeDf, EkJf, EkIg, EfEe, EkJg, EeCg, DfDf, EfCf, DgDf, DfDg, IhIg, EfDe, EkIh, DgCg, DfCg, IbDg, DfEe, FeEf, IbEf, DfEf, IhJf, IhJg, IgIg, EfCh, DgEe, DgEf, EeEg, IhIh, EeDe, IgJg, EkKf, EeCh, IbDf, DgDg, EgDf, FeDg, ElIg, IgIh, DfDe, EjIg, EeCf, DfCh, DgCf, DfCf, DeEe, DkIh, FeDf, EkIf, EeDh, DgCh, IgJf, EjJg, FeEe, DlIh, EgCg, ElIh, EjJf, FeCg, DlIg, IbCg, EfEg, EkJe, FkJf, ElJg, DgDe, DlJg, EgCf, IaEf, FkIg, JaEf, EjIh, EgEf, DkJg, DeEf, EeCi, JgIh, IcEf, EkKe, DkIg, IbEe, EgDg, EeFe, EjKf, IaDf, HhIg, HbDg, ElJf, EfDh, IcDf, EfBh, IcDg, IcCg, FkJg, FeCh, IgKf, FdDg, EkHh, DfDh, DgBh, DfBh, DeDf, DfFe, EfFe, EgEe, EgDe, DkJf, JgJg, IbEg, IbCh, EfBg, DgCe, JlEf, CgCg, HhJf, EeBi, DfBi, IhIf, FeEg, FdEf, EdEf, DlJf, DhCg, JgIg, IeBg, FjIg, FdCh, EdEe, JfIh, JaEe, HhJg, HbEf, HbCh, FkIh, FjJf, ElJe, DhDf, CgDf | All dipeptide pairs which match a combination in the mesostate set are defined as a β-turn. This is based on Rose et. al. (3). Residues classified as turn are included in the coil library. |
| PII-helix | P | Dk, Dl, Ek, El | Residues that have not already been classified as strand, yet match any of the elements in the mesotate set, are categorized as polyproline-II helix. Residues classified as PII are included in the coil library. |
| Coil | C | All | All other residues not classified as P, T, E, or H are assigned as coil. |
Fragment Selection
The secondary structure of all protein residues in the PDB (4) was determined using the method described above. All fragments classified as α helix or β strand were removed. In addition, isolated (one-residue) coil residues were removed, as were residues that lacked all atoms necessary to determine backbone torsion angles. The fragments that remain following this process constitute the Protein Coil Library.
Additional residues are included in the data files that yield information on the context from which the fragment was taken. Whenever possible, the two residues that flank the N-terminal and C-terminal ends of the coil fragment are inlcuded in the database. These four residues are only included if they posess a complete set of backbone atoms; otherwise, they are excluded. As a result, a data files for a four residue fragment will on average contain eight residues: the central four residues represent the fragment itself, and two residues at each end describe the residues flanking the fragment region.
Torsion Angle Calculation
Torsion angles were calculated in the library according to the definitions given here.
References
- Srinivasan, R. and Rose, G. D. "A Physical Basis for Protein Secondary Structure." PNAS 96 (1999): 14258-63.
- Hovmöller, S., et. al. "Conformations of amino acids in proteins." Acta Cryst. D 58 (2002): 768-776.
- Rose, G. D., et. al. "Turns in peptides and proteins." Adv. Prot. Chem. 37 (1985): 1-109.
- Berman, H. M. et. al. "The Protein Data Bank." Nucleic Acids Res. 28 (2000): 235-242.


